Ascertaining when and where genes are expressed is of crucial importance
in order to understand the physiological role of a given gene/protein
and the interactions between them. In addition, the normal expression
patterns can then be compared to those observed in a variety of pathological
conditions to identify pathological hallmarks of gene expression.
The EURExpress, an integrated project funded by the EU under the VI
Framework proposes a transcriptome-wide acquisition of expression
patterns chiefly by means of in situ hybridization (ISH) with non-radioactive
probes and will use this data to establish a web-linked, interactive
digital transcriptome atlas of embryonic mouse. The final goal of
the project is to create the expression data of > 20,000 genes
by RNA in situ hybridization on sagittal sections from E14.5 wild
type murine embryos. This data will result in a detailed description
(at a cellular level) of gene expression patterns in the developing
mouse. The “transcriptome atlas” will be generated using
a newly developed automated RNA in situ hybridization system. Automated
scanning microscopes will collect image data, which will be electronically
sent out in a digital format for annotation. The latter will be performed
using a web-based “virtual” microscope and be entered
in a hierarchical database specifically designed to hold large amounts
of image data and display them in a user-friendly format. For a subset
of genes, mainly those directly involved in human diseases, expression
data will also be generated by using human and murine tissue arrays.
This will offer the opportunity to compare human and mouse expression
patterns in adult tissues. This project builds on a strong European
concentration of skills in gene expression analysis and mouse genomics
and integrates European skills, efforts, resources and information
in the field of systematic gene expression analysis. All expression
data generated by EURExpress will be made readily available to the
scientific community via the EURExpress web-linked database, considerably
advancing our knowledge of gene function and having a significant
impact on the identification of gene expression markers of disease
processes.
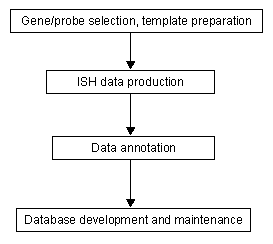
An overall data flow in the project is organised as follows: